Riboswitches
From disordered RNA states to stable tertiary folds – all at single-molecule precision
Bridge the gap between structural dynamics and binding stoichiometry to decode the logic of riboswitches
Riboswitches are segments of mRNA that act as chemical sensors, undergoing structural changes in response to small molecule binding (ions, amino acids, or metabolites) or proteins. This process is fast – it can terminate transcription by RNA polymerase or modulate ribosome binding, inducing a frameshift that produces an entirely new protein from the same transcript. Capturing rapid, dynamic changes and binding kinetics are therefore crucial to understanding and exploiting riboswitch biology.
Observation of riboswitches with single-molecule resolution enables quantification of small molecule binding kinetics and riboswitch conformational changes with high temporal resolution. This allows researchers to understand the interplay between ligand binding, RNA structure, and riboswitch function.
Single-molecule Förster resonance energy transfer (smFRET) captures the real-time dynamics between distinct conformational states, including transient intermediates that remain hidden in ensemble measurements. Fluorescence correlation spectroscopy (FCS) complements this by providing a rigorous quantification of binding affinities and the resulting structural compaction, linking ligand binding to structural changes and function.
Watch riboswitch dynamics unfold with single-molecule resolution
- Resolve conformational transitions, mapping the precise structural changes in riboswitches induced by small molecule binding
- Detect transient states and RNA breathing that are invisible in ensemble assays
- Distinguish conformational selection from induced fit mechanisms upon ligand binding
- Measure dwell times to determine how long a riboswitch stays in a particular conformation
- Identify binding dynamics and kinetics, including cofactor requirements
smFRET is ideal for capturing riboswitch dynamics and ligand-induced conformational changes
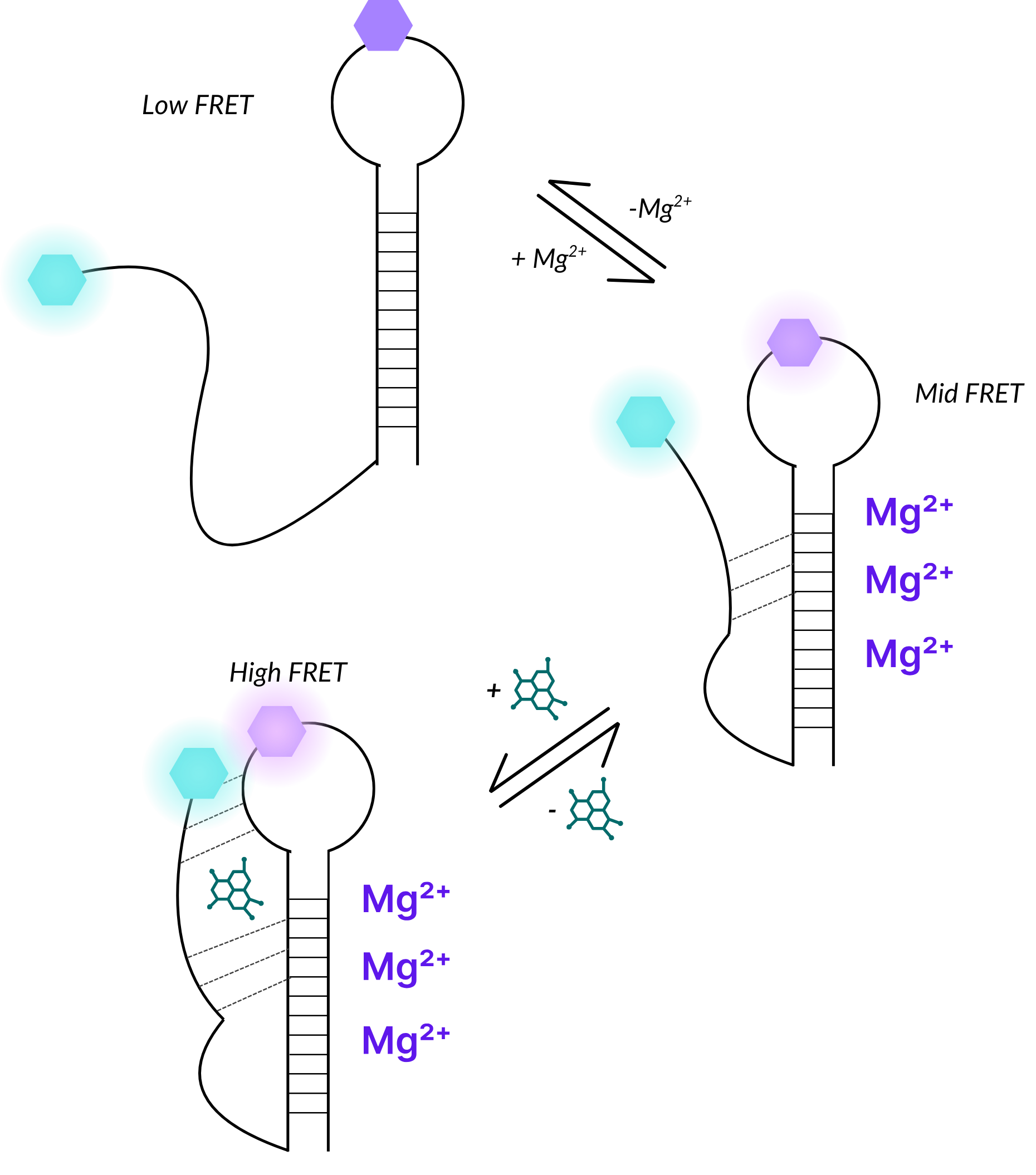
Illustrative example of how riboswitch dynamic structural conformations can be measured using smFRET.
In this example, the unfolded molecule produces a low-FRET state, while the addition of Mg2+ cofactors induces an intermediate conformation with a mid-FRET state, and the binding of a small molecule results in a tightly folded structure with a high-FRET state.
Riboswitch papers and pre-prints featuring EI-FLEX data
Riboswitches FAQs
Why is smFRET a useful tool for studying riboswitches?
smFRET is ideal for identifying transient intermediates and quantifying the conformational dynamics of riboswitches that ensemble assays or static structures would miss.
How can I observe the different functions of the aptamer domain and expression platform?
FCS is ideal for identifying when a ligand has bound the aptamer domain, while smFRET can be used to investigate the ligand-induced structural changes that the expression platform undergoes. Base pairing of the switching sequence to the aptamer domain can also be captured by smFRET.
How does FCS measure the affinity (Kd) of a riboswitch for its ligand?
Can smFRET distinguish between induced fit and conformational selection in riboswitches?
Absolutely. Riboswitches that undergo conformational selection will be highly dynamic and sample a range of conformations, even after ligand binding. Conversely, induced fit riboswitches will undergo a discrete conformational change upon the addition of ligand.
Can I study riboswitches that sense large cofactors?
FCS can indicate whether large cofactors are binding riboswitches; bound riboswitches will diffuse more slowly than unbound molecules. FCCS provides a more direct confirmation of binding as both riboswitch and protein are labelled and their codiffusion is measured, alongside the stoichiometry of the interaction.
How does Mg2+ concentration affect riboswitch measurements?
Can smFRET detect transcriptional pausing?
Riboswitches often work while they are still being synthesised by RNA Polymerase. Using smFRET, you can study co-transcriptional folding by looking at different lengths of the RNA to see at what point the structure becomes competent to bind its ligand and trigger the switch.
How do riboswitch-targeting drugs look in smFRET?
You can use smFRET to see if a drug candidate successfully locks the riboswitch into its inactive state as effectively as the natural ligand does, or if it creates an incorrect structure that the cell can still bypass.
Can I study multi-ligand riboswitches?
Some riboswitches (like the Glycine riboswitch) have two aptamers that work together. Combining smFRET and FCCS is ideal for studying this cooperativity. These techniques in combination can reveal if the binding of the first ligand makes it easier or harder for the second one to bind, or if they must bind simultaneously to trigger the switch.
How does crowding affect riboswitch folding?
In the cell, riboswitches are surrounded by proteins and other RNAs. You can use FCS to measure how molecular crowders, such as PEG, physically compress the riboswitch, often making it more sensitive to its ligand by lowering the energy barrier needed for folding.
"What sets EI-FLEX apart is its versatility - it's our go-to for probing kinetics and conformational changes that the other systems can't fully resolve. This opens up a range of research for us that was previously impossible."
Professor Edwin Antony, Saint Louis University