Antibody-antigen interactions
Resolve the structural fingerprint and binding affinity of antibody recognition
with single-molecule clarity
Capture the binding kinetics of individual antibody-antigen interactions in solution
Characterising antibody-antigen interactions goes beyond just knowing whether binding occurs; it requires understanding the structural stability and binding kinetics under physiological conditions. High-throughput, iterative screening of antibodies and fragments is necessary to assess their functionality during development. Additionally, therapeutic antibodies can undergo structural breathing that may lead to aggregation or loss of potency over time.
There are several assays used to characterise antibody binding affinities. These often average signals, potentially obscuring sub-populations of misfolded antibodies or those with weak affinities. Meanwhile, the required immobilisation on surfaces or plate washing introduces opportunities for experimentally induced artefacts, such as avidity issues. Crucially, these techniques typically do not capture structural changes that ultimately define how well antibodies can bind their targets or induce downstream effects.
Fluorescence correlation spectroscopy (FCS) and single-molecule Förster resonance energy transfer (smFRET) offer a powerful combination for capturing antibody-antigen interactions and the structural changes that influence their functionality and stability.
FCS offers a robust platform for determining antibody binding affinities and kinetic rates, even within complex physiological matrices like serum. Its single-molecule resolution provides the necessary sensitivity to quantify both weak and transient interactions that traditional methods might miss. Complementing this, smFRET enables a deeper look into structural dynamics, capturing the conformational heterogeneity of CDRs and any binding-induced allosteric changes.
Characterise antibody-antigen interactions with single-molecule resolution
- Calculate binding affinities in a freely diffusing environment, avoiding surface-induced denaturation, avidity challenges, and mass-transport artefacts
- Map induced-fit conformational changes to the variable region of the antibody upon antigen binding
- Perform high-precision kinetic measurements directly in physiologically-relevant environments like serum or cell lysates, preserving native protein behaviour
- Identify transient, weak-to-moderate-affinity complexes that can be lost during the mechanical stress of traditional wash steps
- Directly measure the dwell time (koff) of an antibody on its target or inhibition constants (IC50) at the single-molecule level
- Monitor aggregation by simultaneously detecting the presence of aggregates and monomers in a sample
- Quantify complex stoichiometry, definitively counting how many antigens are bound to a single antibody
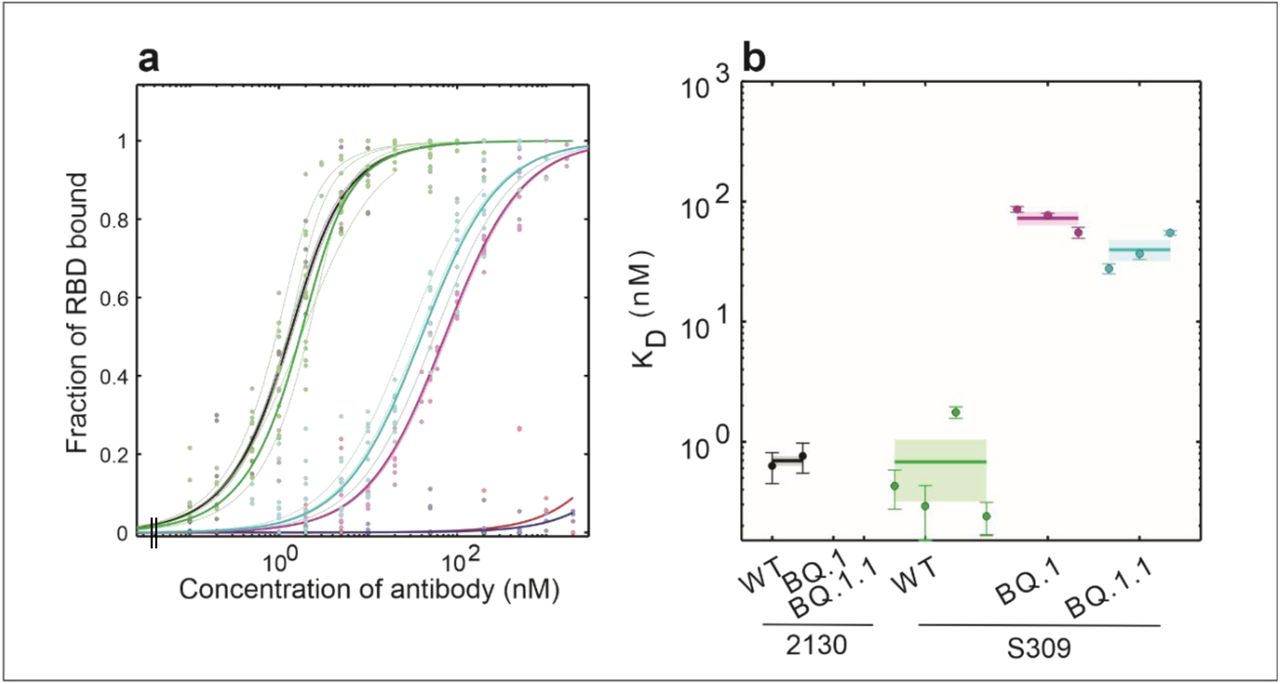
FCS rapidly determined the Kd values for antibody binding against the SARS-CoV-2 spike protein
Kd values were calculated for the binding of two antibodies (wild type and two mutants each) to the receptor-binding domain (RBD) of SARS-CoV-2 spike protein, enabling comparison of how antibody mutations affect affinity.Read the pre-print in full to discover how FCS was performed on the EI-FLEX.
Antibody-antigen papers and pre-prints featuring EI-FLEX data
Antibody-antigen interactions FAQs
How does FCS measure antibody binding affinity (Kd) without washing steps?
Can I determine the stoichiometry of antibody-antigen interactions using FCS?
Using molecular brightness analysis, FCS can count how many labelled antigens are attached to a single antibody. Here, you can clearly distinguish between IgG complexes bound to one or two antigens.
How does FCCS improve the detection of low-affinity antibodies?
FCCS is a far more sensitive technique for low-affinity interactions than standard FCS. By labelling the antibody one colour and the antigen another, co-diffusion can be measured directly. This aids in the differentiation of bound populations from the background noise of unbound molecules, allowing you to detect weak binders that would be missed in other assays.
Can smFRET detect induced fit conformational changes in an antibody?
Many antibodies undergo a hinge movement or a structural shift in the CDR loops upon binding. By placing a FRET pair on the antibody arms, you can see FRET efficiency change the moment the antigen docks.
Can these tools distinguish between specific and cross-reactive binding?
Is it possible to study antibody binding in complex matrices, such as serum or plasma?
FCS is excellent for this because it is a ‘mix-and-read’ technique. You can add your labelled antigen directly to a serum or plasma sample and use the change in diffusion to detect the presence of specific autoantibodies or therapeutic antibodies without needing to purify them first.
How do these tools help in the development of bispecific antibodies?
Bispecific antibodies are designed to bind two different antigens. FCCS is the perfect tool to verify if bispecific antibodies have bound both target antigens. If both targets are labelled with different dyes, a functional bispecific antibody will cause the two colours to cross-correlate, proving it has successfully bridged the two targets.
Can I monitor the stability of antibody-drug conjugates (ADCs)?
For ADCs, the stability of the linker is critical. FCCS is ideal for measuring this. By labelling the antibody and its drug cargo, FCCS can detect if the drug is released prematurely – here the co-diffusion signal would drop to zero. This permits testing of how different pH levels or other factors affect the leakiness of the ADC.
Can I study the binding of nanobodies (VHH fragments)?
Because nanobodies are so small (~15 kDa), their binding to a large antigen doesn’t change the diffusion time enough to be detected by FCS. FCCS is ideal here, because by labelling both the nanobody and its antigen, their cross-correlation can be measured irrespective of molecular weight. smFRET is useful for determining whether binding induces a conformational change in the target protein.
How does pH or ionic strength affect antibody performance in these assays?
FCS can be used to perform a pH-titration to identify under what conditions the antibody is most stable and has the highest affinity. This is critical for optimising antibodies that are intended for use in acidic environments, such as tumours.
"What sets EI-FLEX apart is its versatility - it's our go-to for probing kinetics and conformational changes that the other systems can't fully resolve. This opens up a range of research for us that was previously impossible."
Professor Edwin Antony, Saint Louis University







