Protein-DNA interactions
Observe DNA recognition, regulation, and repair – all with single-molecule clarity
Link structure to function with dynamic data at single-molecule precision
Protein-DNA interactions are highly dynamic. Proteins slide along sequences, bending DNA to bring distal sites together and displacing damaged strands. Many of these functions are multistep processes with transient intermediate states that cannot be observed using ensemble techniques.
By tracking individual molecules in real time, researchers can measure kinetic rates of multistep molecular pathways and map conformational dynamics with high temporal resolution, linking structure to function to drive true mechanistic understanding.
Single-molecule Förster resonance energy transfer (smFRET) and fluorescence correlation spectroscopy (FCS) provide a high-resolution window into how proteins navigate the genomic landscape. While smFRET is ideal for observing conformational changes like DNA bending or helicase unwinding, FCS captures the kinetics and stoichiometry of DNA-protein interactions.
This dual approach allows researchers to observe not just if a protein binds, but how it moves along a sequence, resolves secondary structures, or recruits co-factors to initiate biological signalling, all without the need to isolate individual subspecies within a heterogeneous mixture.
Capture protein-DNA interactions in real time with solution-phase, single-molecule spectroscopy
Observe the molecular events that define recognition, regulation, and repair – one binding event at a time.
- Resolve conformational transitions, mapping the precise structural changes in DNA (e.g., bending, looping, or melting) induced by protein binding
- Observe protein sliding along DNA or the sampling of non-productive conformational states during homology searches
- Distinguish between bound, unbound, and intermediate populations that ensemble methods can’t separate
- Measure the hydrodynamic radius (Rh) of complexes, offering a sensitive assay for studying protein-induced-fit mechanisms or the assembly of large multi-protein complexes on genomic scaffolds.
- Directly observe helicases, polymerases, or nucleases as they move along a template, measuring step size and stall times at the single-molecule level
- Measure biomolecules in solution without tethering, avoiding surface-based artefacts and observing proteins that cannot be immobilised
- Quantify dynamically interconverting nanoscale distances with Å precision
- Pre-configured smFRET and FCS methods streamline setup so you can focus on discovery
- Generate reproducible, high-quality data that moves seamlessly from acquisition to publication
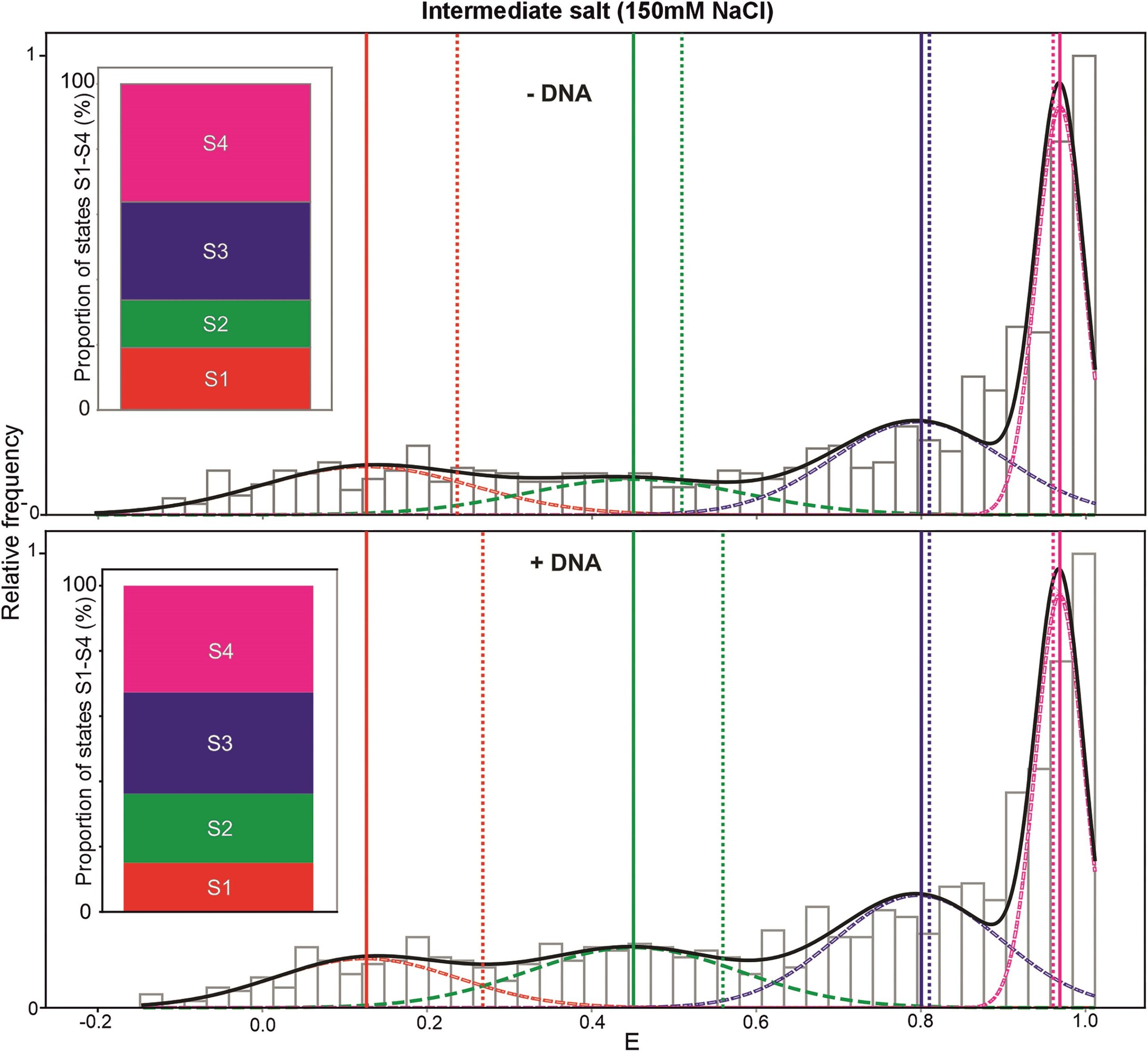
smFRET reveals two previously unknown intermediate states of the bacteria Rep helicase
In a recent publication, researchers used smFRET to explore the conformational states of the bacterial Rep helicase. The open (S1) and closed (S4) conformations had previously been identified in crystal structures – smFRET revealed two additional, undiscovered conformational states (S2 and S3). The addition of DNA alters the proportion of Rep molecules in all states; the presence of DNA increases the percentage of molecules in the S2 state and reduces those in the S1 state
Read the paper in full to discover how smFRET was performed on the EI-FLEX.
Protein-DNA papers and pre-prints featuring EI-FLEX data
Protein-DNA interactions FAQs
What protein-DNA interactions can I study?
In theory, the EI-FLEX can study almost any protein-DNA interaction, although the choice of labels and their positions should be carefully considered to preserve biomolecular function and structure. Examples from EI-FLEX customers include polymerases, retroviral integrases, and interactions of single-stranded DNA with replication proteins.
Can I distinguish on- vs off-target binding of sequence-dependent DNA-binding proteins?
FCS is used for quantifying binding affinity. On-target binding of DNA-binding proteins typically exhibits significantly lower Kd values than those that are off-target. By performing a titration, you can distinguish these populations based on the concentration-dependent shift in diffusion dynamics.
smFRET can complement this by providing high-resolution insights into conformational changes upon binding. Specific interactions typically yield stable, discrete FRET states corresponding to defined structural architectures. Conversely, non-specific binding is often more transient and dynamic; these interactions can be identified using Burst Variance Analysis (BVA) of smFRET data to detect rapid interconversion, which appears as increased variance compared to the shot-noise limit.
Can I detect multiple proteins binding to DNA and the stoichiometry of DNA-protein complexes?
If the same protein binds DNA multiple times, FCS can be used to measure brightness changes, which correlate with the number of bound proteins.
FCCS determines whether two different types of proteins are bound to the same DNA molecule in solution and can calculate the stoichiometry of complexes.
How can I measure exonuclease or endonuclease activity?
FCCS is ideal here as it can be used to observe a loss in co-diffusion upon cleavage if both DNA strands are labelled; FCS can complement this by capturing a change in molecular size as DNA is fragmented.
smFRET can be used to monitor either DNA or protein conformational state and dynamics during nuclease activity.
Can I detect strand invasion or displacement events?
smFRET can detect strand displacement – by labelling the two original strands, FRET efficiency will decrease and eventually drop to zero as the invading strand induces displacement. This is a very dynamic process, and burst variance analysis and Hidden Markov modelling of smFRET data are ideal to capture branch migration and the discrete steps of strand invasion.
FCCS can also confirm that displacement has been successful by labelling the invading strand – an increase in cross-correlation confirms that displacement has occurred.
Can I study protein binding to supercoiled DNA?
smFRET is ideal for characterising how binding of proteins such as topoisomerases alters the architecture of supercoiled DNA. For example, dye pairs might be positioned such that they are in close proximity on supercoiled DNA and move apart upon protein-induced relaxation. This would cause a measurable change in FRET efficiency, from high to low.
Meanwhile, FCS can detect the changes in hydrodynamic radius as supercoiled DNA relaxes, shifting from a tightly compacted molecule to one that has a looser structure.
Can I determine if DNA length influences binding kinetics?
What are the minimum and maximum lengths of DNA that I can use?
Two factors are important for considering the length of DNA used for solution-based FCS and smFRET. For FCS, the DNA is ideally smaller than the diameter of the confocal volume so that it may. For smFRET, the distance between the donor and acceptor fluorophores must be within the dynamic range of FRET detection (between 1 and 100 angstroms).
Exceptionally small DNA fragments run the risk of having their dye pairs interact with each other. Additionally, if the DNA is shorter than the Förster radius of the chosen dye pair (typically around 5-6 nm or 15-18 base pairs), FRET efficiency will be perpetually at 100 %. At the upper end, DNA molecules that are very long run the risk of having dyes placed further than 10 nm apart, meaning that FRET will never occur.
Can I study the effects of buffer conditions, such as pH and ionic strength, on DNA-protein interactions?
Absolutely, although it should be noted that pH and salt ions can affect fluorescent dyes and the structure of DNA, irrespective of whether a protein is bound. The refractive index of buffers can also change with the addition of salt, requiring recalibration and potentially recalculation of Förster radii.
Can I detect protein-induced conformational changes in DNA, such as looping or bending?
smFRET is ideal for detecting whether looping or bending of DNA is caused by protein binding, or whether free DNA is undergoing ‘breathing’; these are often distinguished as a discrete, static change and dynamic interconversions, respectively.
"What sets EI-FLEX apart is its versatility - it's our go-to for probing kinetics and conformational changes that the other systems can't fully resolve. This opens up a range of research for us that was previously impossible."
Professor Edwin Antony, Saint Louis University