Enzymes
Resolve the conformational landscape of catalysis and enzyme inhibition with single-molecule clarity
Visualise catalytic cycles and determine the structural dynamics of enzyme inhibition
Enzymes are the biological catalysts that sustain life, yet their function is rarely the simple ‘lock and key’ mechanism. Most enzymes are dynamic machines that rely on rapid structural fluctuations to capture substrates and release products. Understanding these dynamics is the key to mastering metabolic regulation, designing high-efficiency biocatalysts, and developing both competitive inhibitors and allosteric drugs.
To understand this, we must look beyond the average. Single-molecule insights are vital for capturing the transient states and short-lived intermediates that are typically masked by ensemble-averaging techniques.
Single-molecule Förster resonance energy transfer (smFRET) allows the visualisation of how enzymes move through catalytic cycles, undergoing dramatic conformational changes as substrates bind and are turned over. This single-molecule resolution can capture transient, high-energy structural intermediates that occur during the millisecond window of a chemical reaction.
Fluorescence correlation spectroscopy (FCS), provides additional insights by determining precise kinetic parameters for substrate binding and stoichiometries of multi-subunit complexes.
Observe individual enzymes and reveal hidden intermediates with rapid, in-solution dynamics
- Map the structural response of active sites and distal domains to competitive or allosteric inhibitors
- Detect short-lived, high-energy conformational intermediates that are invisible to bulk methods
- Detect heterogeneity within enzyme populations to understand why some are more efficient than others
- Distinguish between induced-fit and conformational selection mechanisms
- Identify the oligomeric state of monomers and multimers, linking assembly to activity
- Calculate binding dynamics and kinetics by analysing the fluctuations in fluorescence as substrates are converted to products
smFRET captures global conformational changes in Rag GTPase upon guanine nucleotide binding
In the absence of guanine nucleotides, Rag GTPase occupies an open, dynamic conformation (low FRET efficiency). When both sites are occupied by GDP, Rag GTPase remains in an open, static conformation (low FRET efficiency). When both GDP and GTP are bound, the protein predominantly adopts a static closed conformation.
Read the paper in full for more information on how smFRET was performed on the EI-FLEX.
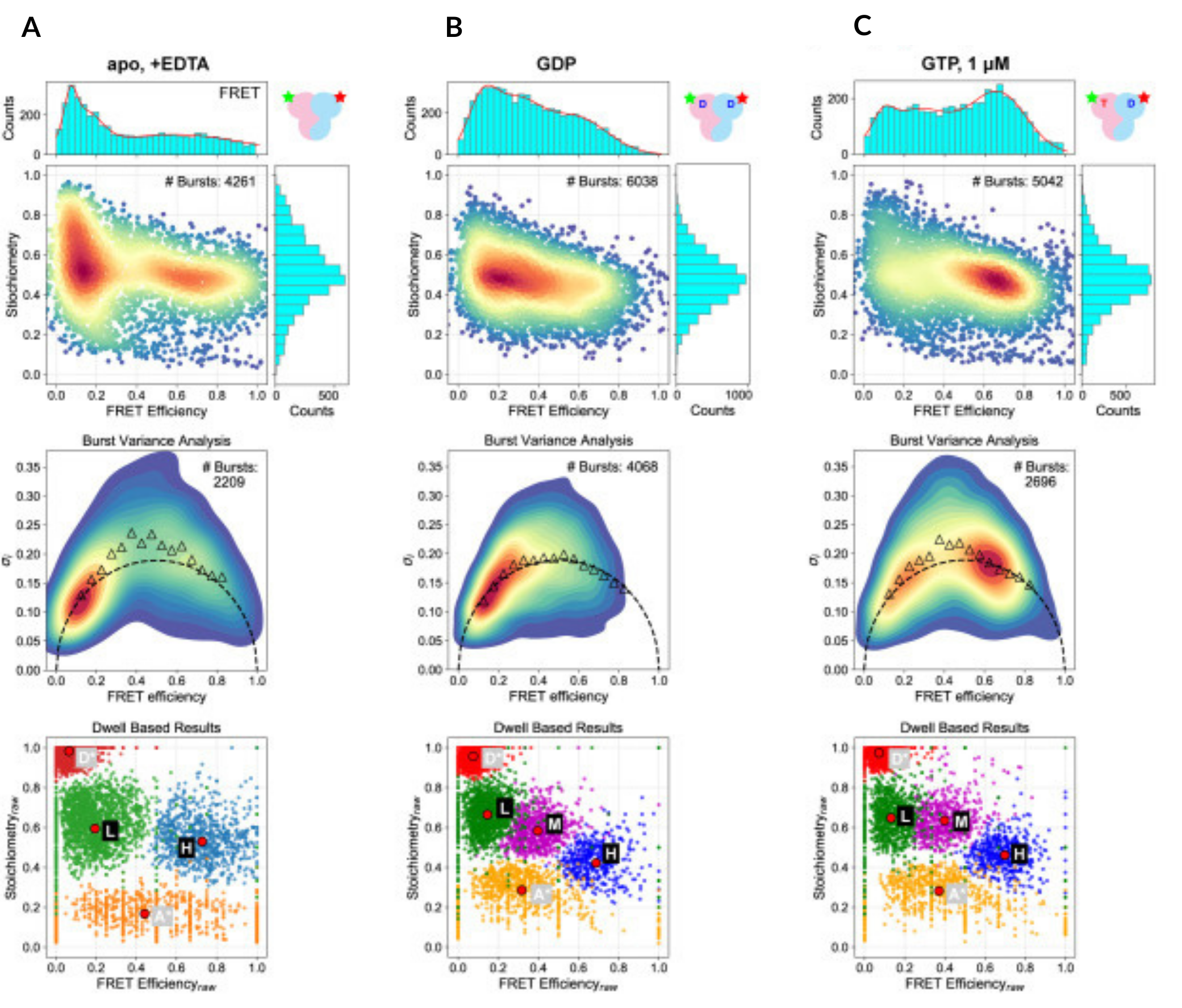
A) Unbound Rag GTPase (EDTA only)
B) Rag GTPase with GDP only
C) Rag GTPase with GDP and GTP
Figure taken from: Doxsey, D. D. & Shen, K. Global conformation of the Rag GTPase heterodimer governs eukaryotic amino acid sensing. Proceedings of the National Academy of Sciences 122, e2517050122 (2025)
Enzymes papers and pre-prints featuring EI-FLEX data
Enzymes FAQs
How does smFRET monitor the conformational cycle of an enzyme?
By placing a donor and acceptor dye on two moving domains, changes in FRET efficiency can be measured as the enzyme captures a substrate, performs catalysis, and releases the product.
How does FCS measure enzyme-substrate affinity (Km)?
Can FCCS prove that an enzyme and its cofactor are interacting?
By labelling both the enzyme and cofactor, their cross-correlation signal confirms that they have formed a functional holoenzyme complex. The action of inhibitors can also be quantified; the cross-correlation signal disappears with successful inhibition.
Can smFRET detect allosteric changes?
smFRET is an ideal technique for visualising structural changes in an active site induced by molecule binding at a distant site, provided that the changes are within the dynamic range of FRET.
Can I see the difference between induced fit and conformational selection?
By analysing FRET efficiencies before and during binding, the specific mechanism can be determined by smFRET. During conformational selection, enzymes interconvert between conformations, even in the presence of ligands. For induced-fit mechanisms, substrate binding forces the enzyme to change shape – Doxsey and Shen demonstrated this in their recent paper exploring the conformational changes of Rag GTPase.
Can I study enzymes that undergo multiple rounds of catalysis?
For enzymes that stay attached to a substrate, smFRET can monitor how the enzyme transitions through conformational states over the course of multiple rounds of catalysis.
Can FCS detect enzyme inhibition?
FCS can detect inhibition by competitive and allosteric inhibition. By performing a titration of an inhibitor, a greater amount of labelled substrate will be prevented from binding, which will result in faster diffusion through the confocal volume compared to substrates bound by enzymes. Some inhibitors cause the enzyme to aggregate or denature, which FCS detects as a massive shift in brightness alongside a reduced rate of diffusion.
Can smFRET observe product inhibition?
If the product of a reaction stays stuck in the active site, this inhibition can be captured by smFRET by observing stalled conformational dynamics. Because single enzymes are observed, stalled molecules can be differentiated from those that have slower enzymatic cycles.
How does molecular crowding affect FCS measurements of enzyme kinetics?
FCS is often used to measure the diffusion of an enzyme in a crowded mimic buffer, such as PEG or Ficoll, although this can stabilise the active conformation of the enzyme, making it more efficient than it appears in a standard, dilute lab buffer. FCS can also be performed in complex matrices such as serum and cell lysates.
“Solution-based measurements, like those taken using the EI-FLEX, avoid surface tethering, which can introduce artefacts, especially for proteins that don’t behave well near hydrophobic surfaces. Keeping proteins in solution lets you observe their native conformation, and it’s much easier experimentally: you label, dilute, and measure, often getting results in minutes rather than days or weeks with TIRF-based systems."
Associate Professor Kuang Shen, UMass Chan





