Motor proteins
Observe motor protein conformational changes and cargo binding that drive cellular transport
Capture the structural heterogeneity of motor proteins with single-molecule clarity
Motor proteins are the workhorses of the cell, driving essential biological processes through coordinated, large-scale conformational changes. While they are known to walk along cytoskeletal filaments to deliver cargo, the finer mechanics of their regulation – and the molecular roots of their dysfunction in disease – remain elusive.
To understand this, we must look beyond the average. Single-molecule insights are vital for capturing the transient states and short-lived intermediates that are typically masked by ensemble-averaging techniques.
By integrating single-molecule Förster resonance energy transfer (smFRET) and Fluorescence correlation spectroscopy (FCS), we can complement high-resolution static structures with real-time dynamic data. This approach maps complex stoichiometry and kinetics, providing the missing link between motor protein architecture and physiological function.
Bridge the gap between motor protein structure and complex cellular traffic
- Monitor the conformational changes within the motor’s lever arm, linking ATP consumption directly to physical motion
- Measure the exact stoichiometry of how many motors are engaged in a transport complex
- Quantify the dynamics of cargo binding and transport, from binding affinity to cargo specificity
- Characterise the structure of kinesin-binding proteins and the mechanisms of trans-inhibition that stall transport
- Identify the specific conformational triggers that transition a motor from an autoinhibited state to active motility
smFRET can distinguish highly heterogeneous Kinesin-1 populations
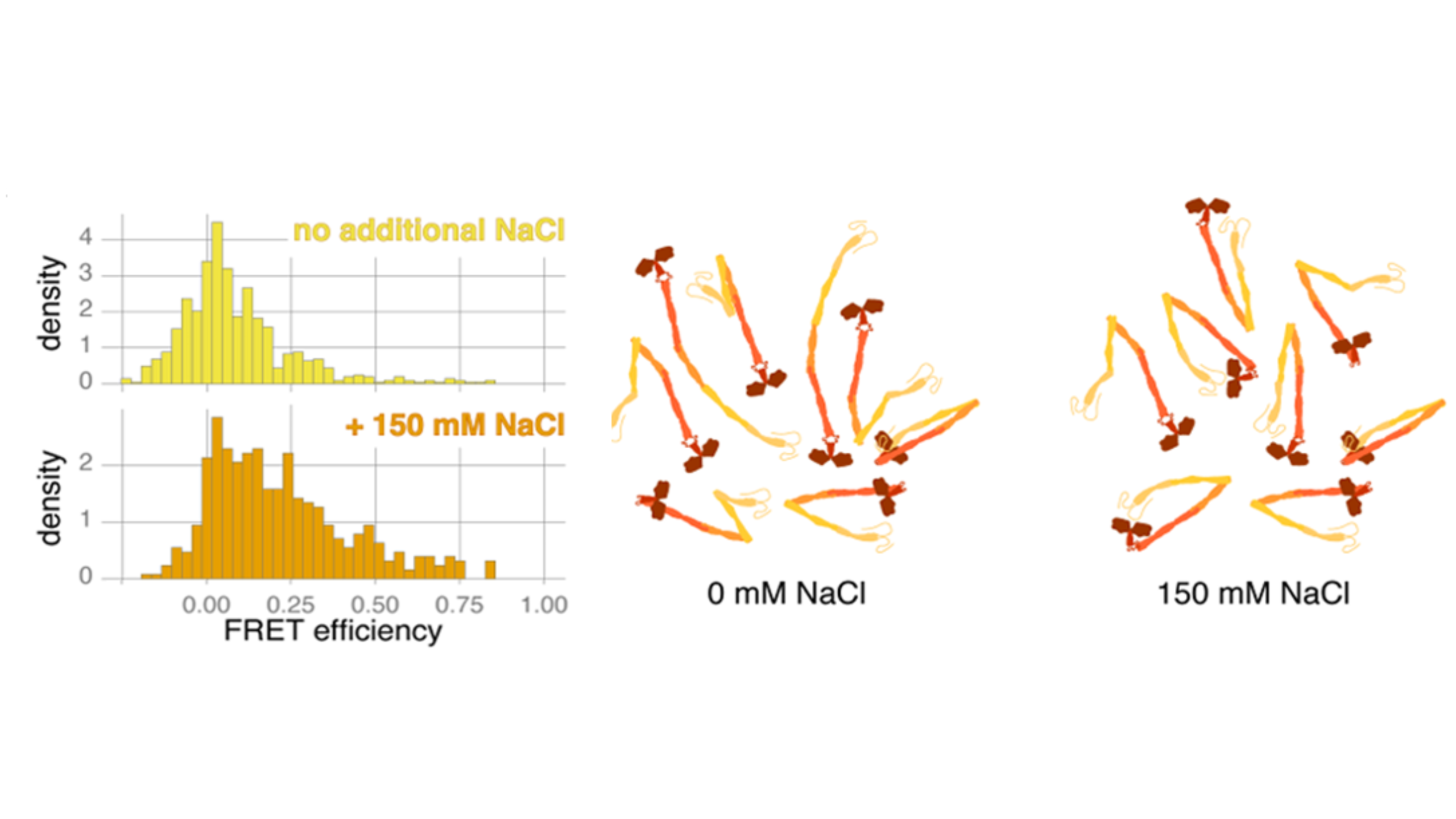
Smith et al. used smFRET to investigate the impact of salt on the structural conformation of Kinesin-1. At 0 nM NaCl, Kinesin-1 is thought to have a mixed population that is mostly formed of molecules in open conformations (low FRET efficiency). At 150 nM NaCl, a larger proportion of these molecules are in a closed, inactive state (high FRET efficiency), although both conditions show highly heterogeneous populations. If this were an ensemble assay, the FRET efficiency from multiple conformations would be averaged together, resulting in a single FRET value that would suggest the presence of a single conformation. smFRET allows you to see each kinesin-1 molecule and its conformational state, revealing the true heterogeneity of these molecules.
See the paper on this study to explore how smFRET was performed on the EI-FLEX.
Motor proteins papers and pre-prints featuring EI-FLEX data
Motor proteins FAQs
Can I observe the power stroke of a motor protein?
By placing a FRET pair between the motor domain and the lever arm, smFRET can detect conformational changes triggered by ATP hydrolysis.
How can I confirm if a motor protein is carrying a specific cargo?
If the size discrepancy between the motor and the cargo is large enough, FCS can be used to observe binding by labelling the smaller of the two molecules. Alternatively, if both the cargo and the motor can be labelled, FCCS can be used to monitor binding by observing coincident photon bursts from each labelled molecule as the complex moves through the confocal volume.
How do I study the binding of multiple motors on one cargo?
Using an FCS brightness analysis, you can measure the number of motor proteins attached to a single transport vesicle. Additionally, FCCS can identify whether two different motor proteins are binding to the same cargo together.
Can smFRET resolve the different states of the ATP hydrolysis cycle?
Motor proteins change shape depending on what is in their active site. smFRET can assign a specific conformational fingerprint to each chemical state.
Can I study kinesin autoinhibition?
smFRET can identify conformational changes between folded (inactive) and unfolded (active) states of kinesins. In a recent publication, researchers used the EI-FLEX to measure how salt concentration affects Kinesin-1 autoinhibition – smFRET was ideal here because it could capture how heterogeneous and dynamic Kinesin-1 structures are, even in the presence of salt.
Can I monitor a motor protein remodelling a protein complex?
For motors like p97 or ClpB, which use ATP to unfold other proteins, you can use smFRET to label the substrate protein. You can watch in real-time as the motor pulls on the substrate, causing it to unfold, resulting in a change in FRET efficiency.
“What sets the EI-FLEX apart is its versatility - it's our go-to for probing kinetics and conformational changes that the other systems can't resolve. This opens up a range of research for us that was previously impossible.”
Professor, Saint Louis University