G-quadruplexes and other higher-order structures
Unravel the complexity of higher-order DNA structures one molecule at a time
Observe real-time folding dynamics of higher-order structures with in-solution fluorescence spectroscopy
DNA can undertake a range of conformations beyond the classic B-DNA helix. Higher-order structures are of great interest for understanding crucial biological processes, but capturing their formation is challenging due to their dynamic, transient nature. They are highly sensitive to environmental changes, and pathways may be formed of short-lived intermediates.
Single-molecule Förster resonance energy transfer (smFRET) and Fluorescence Correlation Spectroscopy (FCS) are ideal for dissecting the folding pathways and topological transitions of higher-order nucleic acid structures, adding complementary data to structural and biophysical workflows.
smFRET serves as a sensitive probe for monitoring the proximity of specific nucleotides, enabling the detection of transitions between linear, looped, and multi-stranded geometries. FCS quantifies the individual and co-diffusion of distinct nucleic acids alongside the hydrodynamic changes associated with the transition from disordered single strands to highly compacted tertiary structures.
Visualise dynamic higher-order structures with single-molecule clarity
Bring higher-order structures to life by augmenting your structural biology toolkit.
- Explore the full range of higher-order conformations in solution, without immobilisation, fixation, or crystallisation
- Observe folding intermediates and quantify the kinetic barriers involved in the formation of triplexes, i-motifs, and pseudoknots
- Perform stoichiometric analysis of multi-strand assemblies
- Observe 'breathing' dynamics or transient base-pair fraying under physiological conditions that are invisible in ensemble methods
- Resolve coexisting topological isoforms, such as the parallel, anti-parallel, and hybrid states of G-quadruplexes
- Titrate metal ion-dependent folding and identify the conditions required for the formation of compact, functional tertiary structures
smFRET can distinguish between G-quadruplex topological isoforms
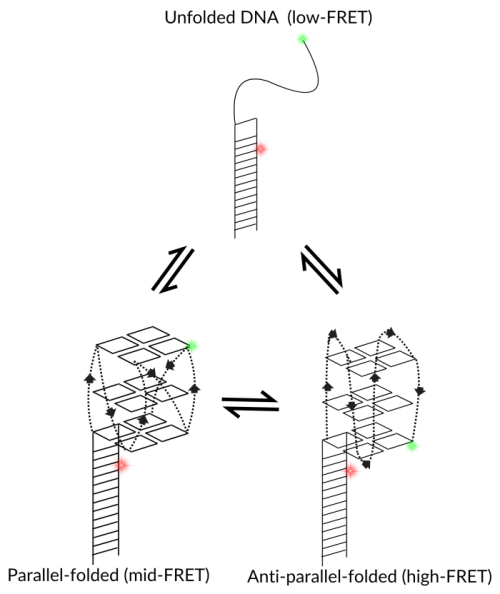
Folding of G-quadruplexes can be captured using single-molecule spectroscopy, determining the differences between parallel and anti-parallel structures. In the illustrated example, the placement of fluorescent dyes leads to low, mid and high-FRET efficiencies depending on particular folding conformations.
Ternary complex papers and
pre-prints featuring EI-FLEX data
Protein structure and dynamics FAQs
How does smFRET visualise a G-quadruplex?
By placing a dye pair at the ends of the guanine-rich sequence, smFRET can be used to monitor the folding of a G-quadruplex. A linear strand shows a low FRET state (dyes are far apart) with a likely broad distribution of FRET efficiency, while the folded structure pulls the dyes into close proximity, resulting in a distinct, high FRET state.
Can smFRET distinguish between different G-quadruplex topologies?
G-quadruplexes can fold into parallel, anti-parallel, or hybrid topologies depending on the salt ions present. Each topology has a slightly different compact shape. smFRET can resolve these as different FRET efficiencies, as shown in the figure above.
How does FCS measure G-quadruplex stability?
FCS measures the overall compaction of DNA. A folded G-quadruplex is much more compact than a random-coil single strand. By measuring the diffusion time, you can calculate the melting temperature and stability of G-quadruplexes, even at low concentrations. smFRET can also be used to measure stability given that it is an ideal tool for measuring conformational dynamics.
Can smFRET monitor the folding of an i-motif?
i-motifs are cytosine-rich structures that fold at acidic pH. By performing a pH titration while monitoring smFRET, you can watch the DNA fold into an i-motif as the pH drops.
Can I study proteins that unfold quadruplexes?
Many helicases and specialised G4-resolvases bind and unwind these structures. By separately labelling the G-quadruplex and the protein, FCCS can directly confirm whether the protein has bound (both signals will codiffuse). smFRET can complement this by providing structural information; FRET efficiency will change as the protein unfolds the quadruplex.
Can these tools detect G-quadruplex-ligand interactions for drug discovery?
Small molecules, such as G-quadruplex-stabilisers, are being developed as anti-cancer drugs. Using smFRET, you can determine if a drug candidate successfully locks the G-quadruplex in a folded state, even under conditions where it would normally unfold (like low salt or high temperature).
Can I measure the folding kinetics of G-quadruplexes using salt?
Using smFRET, folding kinetics can be measured by titrating salt concentrations and observing the rates of G-quadruplex folding. Different ions stabilise G-quadruplexes to varying extents, which can also be captured using this technique.
Can smFRET detect alternative structures like triplexes or H-DNA?
In theory, any structure that involves a distance change can be studied using smFRET; the formation of triplex DNA would result in a distinct FRET efficiency from that of the duplex baseline. This allows you to study the stability and formation of H-DNA in supercoiled regions. FCCS can also confirm that these two structures have hybridised.
Can I see the structural polymorphism of a single G-quadruplex sequence?
smFRET captures individual G-quadruplexes, visualising heterogeneous populations that would likely be lost in ensemble data. A single molecule may flicker between two FRET states, indicating that it is jumping between two G-quadruplex topologies at equilibrium.
"What sets the EI-FLEX apart is its versatility - it's our go-to for probing kinetics and conformational changes that the other systems can't resolve. This opens up a range of research for us that was previously impossible."
Professor Edwin Antony, St. Louis University






