In this application note, we demonstrate how fluorescence cross-correlation spectroscopy (FCCS) can be performed on the EI-FLEX Pro system to characterise the formation of a ternary complex. A model DNA oligo system was used for the purposes of this application note, although FCCS is also ideal for studying other ternary complexes, such as PROTACs, molecular glues, and bispecific antibodies.
Here, direct quantification of ternary complex formation was performed using FCCS. We generated hook plots to establish the optimal concentration of the unlabelled oligo to produce the desired stoichiometry and avoid unwanted dimers. The impact of salt concentration on hybridisation kinetics was also investigated.
Overview of this application note:
- FCCS provides a direct quantification of ternary complexes in solution, as cross-correlation is only detected when all three components are present in the same complex
- Hook plots produced by titration of the unlabelled oligo identify the optimal concentration for ternary complex formation
- Kinetics of ternary complex formation can be directly calculated using FCCS
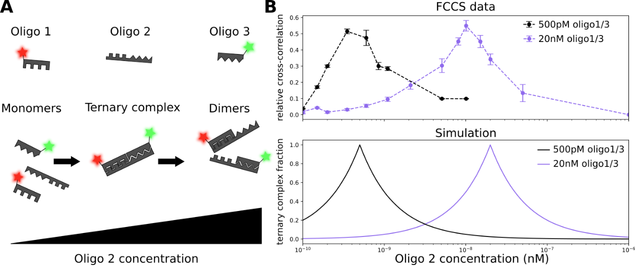
Figure 1 – Hook plots generated using FCCS analysis of ternary complexes
(A) Schematic of the dose-response to the ternary-complex inducing oligo 2. With increasing concentration of oligo 2, a ternary complex is formed; at saturating concentrations, only dimers of oligo 1/3 and oligo 2 are formed.
(B) Top: Dose-response curve at 500pM (black) and 20nM (purple) oligo 1/3.
Bottom: Calculated dose-response curves using the model developed by Douglass et al.



