Protein-RNA interactions
Capture ternary structure formation and dynamic regulation at single-molecule precision
Explore RNA recognition, regulation and remodelling with single-molecule spectroscopy
RNA molecules form complex structures that are highly influenced by protein binding. From the ribosome to the spliceosome, RNA remodelling and degradation via interaction with proteins are vital steps in cellular control. These processes are highly dynamic, meaning static or ensemble methods may obscure the true flexibility and range of conformations that RNA can take.
By tracking individual molecules in real time, researchers can measure complex kinetics, map dynamic conformational shifts with high temporal resolution and quantify binding and complex formation, linking structure to function to drive true mechanistic understanding.
Single-molecule Förster resonance energy transfer (smFRET) and fluorescence correlation spectroscopy (FCS) provide a single-molecule view of how proteins navigate the complex transcriptomic landscape. While smFRET captures the nuances of RNA tertiary folding, the remodelling of intricate hairpins, and the structural remodelling driven by helicases, FCS quantifies the absolute binding affinity and stoichiometry of complexes in solution.
This dual approach allows researchers to observe not just if a protein binds, but how it chaperones RNA conformation, resolves inhibitory secondary structures, or recruits co-factors to coordinate post-transcriptional regulation.
Capture protein-RNA interactions with in-solution, single-molecule clarity
Observe individual molecular events that define recognition, regulation and remodelling to unravel the complexity of protein-RNA interactions.
- Map the precise RNA structural changes induced by protein binding with Å precision, such as riboswitch flipping
- Define the dynamic interconversions, kinetics and stoichiometry of protein interactions with RNA, such as RNases, ribosomes and small nuclear ribonucleoproteins
- Distinguish bound, unbound and intermediate populations that ensemble methods can’t separate
- Measure protein-RNA interactions in solution without tethering, avoiding surface-based artefacts
- Distinguish between heterogeneous, static populations and dynamically interconverting species
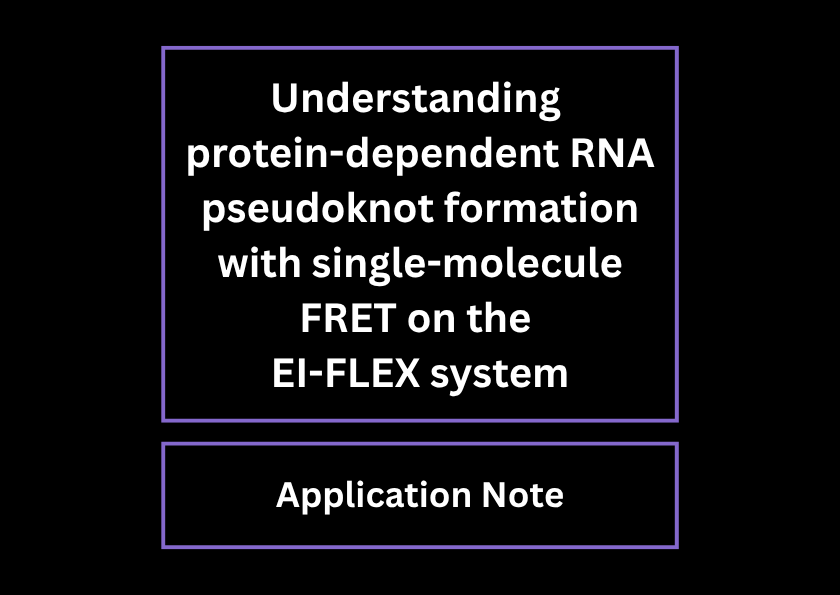
Understanding protein-dependent pseudoknot formation using smFRET
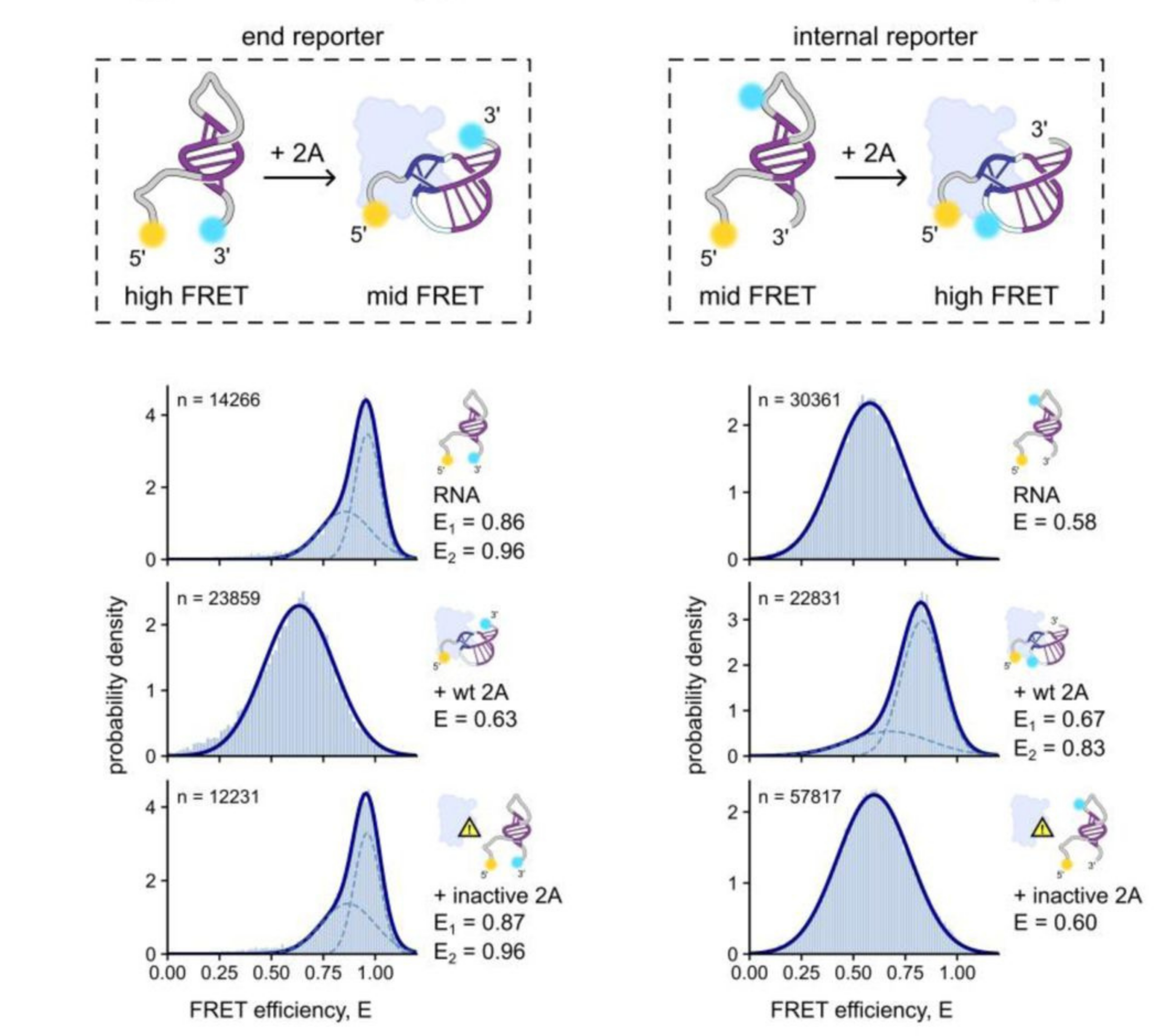
In a recent paper, smFRET added complementary information to structural techniques such as X-ray crystallography and SAXS to elucidate RNA conformational changes upon protein binding. The authors used dye pairs in two places (as an end reporter and internal reporter) to distinguish stem loop from pseudoknot structures.
Read the pre-print in full to discover how smFRET was performed on the EI-FLEX.
Protein-RNA papers and pre-prints featuring EI-FLEX data
Protein-RNA interactions FAQs
What types of protein-RNA interactions can be studied with the EI-FLEX?
In theory, so long as the position of the fluorescent dyes doesn’t disrupt biomolecular function, the EI-FLEX can study almost any protein-RNA interaction. Examples from EI-FLEX customers include RNA-dependent RNA polymerases and pseudoknots.
How does smFRET distinguish between protein-binding-induced structural changes and RNA secondary structure folding?
smFRET can differentiate between RNA secondary structures, such as hairpins or pseudoknots that form in the absence of protein, and protein-induced conformational changes. To achieve this, smFRET should be performed for RNA alone and in the presence of the binding protein.
Can I measure the binding affinity (Kd) of protein-RNA interactions?
Can I use smFRET to detect conformational changes in riboswitches upon ligand binding?
Absolutely – smFRET can monitor how RNA structure shifts in response to a small molecule ligand and how these structural changes are influenced by environmental factors such as salinity.
Can these techniques monitor the assembly of large Ribonucleoprotein (RNP) complexes?
If the RNP is large enough, FCS can be used to detect whether a labelled protein is interacting with it. By labelling two protein components with different colours, FCCS can monitor their interaction with an RNP complex within the same experiment.
Can I detect RNase activity in real-time?
In FCS, RNA degradation leads to a rapid decrease in diffusion time as the labelled fragments become smaller and faster. FCCS can also be used here to capture a loss in codiffusion as RNA is degraded.
How does ionic strength (e.g., Mg2+) affect RNA-protein measurements?
RNA folding is extremely dependent on divalent cations, meaning cation-induced RNA folding and protein-induced stabilisation. Both smFRET and FCS require careful RNA-only buffer controls for every salt concentration used.
What should I consider when labelling RNA for RNA-protein studies?
RNA can be sensitive to phototoxicity. Therefore, additives like GODCAT, PCA/PCD, and trolox are often added to scavenge oxygen species that can damage RNA. Fluorophores are also vulnerable to quenching from proximity to guanine bases, resulting in dim signals. Overall, ease of labelling, preservation of function, and compatibility with the optical properties of the instrumentation are important considerations when choosing fluorophores.
How do I determine the stoichiometry of multiple proteins binding to a single mRNA?
FCS can determine the number of singly-labelled proteins within a complex in the confocal volume. FCCS expands this by labelling two proteins separately to see if they both bind mRNA.
"What sets EI-FLEX apart is its versatility - it's our go-to for probing kinetics and conformational changes that the other systems can't fully resolve. This opens up a range of research for us that was previously impossible."
Professor Edwin Antony, Saint Louis University







