RNA Structure
Decoding the structural diversity of the RNA world
Resolve complex RNA folding, tertiary motifs, and transient intermediates at single-molecule resolution
RNA is a highly dynamic molecule, folding into complex architectures that define its function. From hairpins and stem loops to complex tertiary structures such as A-minor motifs and ribozymes, these molecules experience a wide range of conformations and interconversions, despite RNA having a relatively transient lifetime compared to other biomolecules.
Due to their flexibility, RNA populations can be heterogeneous, meaning distinct species or those present at low abundance may be lost in ensemble measurements. Like proteins, RNAs also pass through transient indeterminates that only exist for microseconds, requiring techniques with high temporal resolution to capture these short-lived conformations.
Single-molecule Förster resonance energy transfer (smFRET) maps the transitions between RNA conformational states, exposing the short-lived folding intermediates that govern function. Fluorescence correlation spectroscopy (FCS) complements this by quantifying RNA interactions with other biomolecules, such as the binding of small molecules or proteins, and characterising how this impacts structural compaction.
Map dynamic RNA folding landscapes and biomolecular binding
- Identify folding intermediates or misfolded structures
- Quantify the stability of pseudoknots and long-range interactions that dictate RNA stability and translation
- Measure the stoichiometry and binding affinity of RNA-binding proteins (RBPs) as they remodel RNA structure
- Titrate pH or salt ions to observe how RNA scaffolds respond to environmental stressors
- Resolve structural switches in viral genomes that allow the virus to alternate between translation, replication, and packaging
- Track the subtle movements and aminoacyl-stacking dynamics of tRNA
- Quantify the binding kinetics of small regulatory RNAs to their targets and observe the resulting structural changes in the 3' UTR or seed region
Understanding protein-dependent pseudoknot formation using smFRET
In a recent paper, smFRET added complementary information to structural techniques such as X-ray crystallography and SAXS to elucidate RNA conformational changes upon protein binding. The authors used dye pairs in two places (as an end reporter and internal reporter) to distinguish stem loops from pseudoknot structures.
Read the pre-print in full to discover how smFRET was performed on the EI-FLEX.
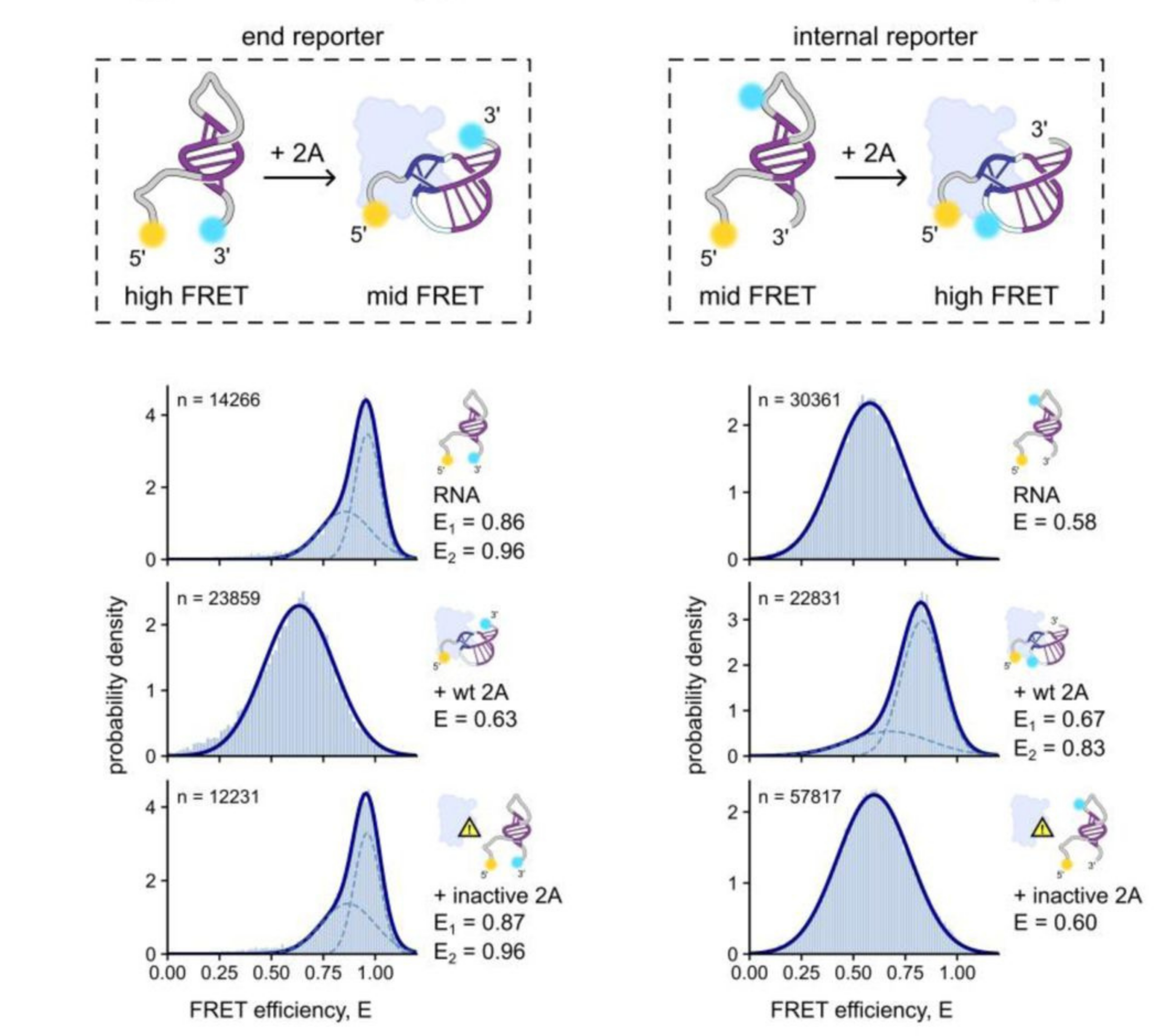
Schematics of dye placement and conformational changes (above). Corrected FRET histograms with Gaussian fits for RNA only, RNA + wildtype protein 2A, and RNA + inactive protein 2A (below)
Left) End reporter dye pair
Right) Internal reporter dye pair
Figure taken from: Betts, J. K. et al. A new protein-dependent riboswitch activates ribosomal frameshifting. bioRxiv 2025.07.17.665365 (2025) doi:10.1101/2025.07.17.665365.
RNA structure papers and pre-prints featuring EI-FLEX data
RNA structure FAQs
How does smFRET on the EI-FLEX differ from chemical probing methods like SHAPE-Seq or DMS-MaPseq?
Chemical probing provides a static, ensemble-averaged map of which nucleotides are paired or unpaired. In contrast, smFRET provides the dynamic distance between specific points. FRET allows you to observe structures that flicker between states and identify transient intermediates that are invisible to sequencing-based methods.
Can I study very large RNA molecules, such as lncRNAs?
Absolutely. By placing fluorophores at key junctions or distant domains, you can monitor the global folding or the local breathing of specific architectural motifs within a massive lncRNA scaffold.
Is it possible to observe RNA-protein interactions in real-time?
Can I measure the effects of different ions on folding?
Yes – RNA folding is highly dependent on the ionic environment. smFRET is ideal for observing how titrations of divalent cations or monovalent salts influence the conformational dynamics of RNA structures.
Can smFRET resolve multiple folding states in a single sample?
This is a core strength of the technique. Instead of one average value, the data are displayed as a histogram of FRET efficiencies. If your RNA exists in three different conformations (e.g., unfolded, hairpin, and pseudoknot), you will see three distinct peaks, allowing you to quantify the population percentage of each.
Do I need to immobilise my RNA to get high-quality structural data?
No – smFRET and FCS observe RNA in a freely diffusing, native-like state, avoiding any structural artefacts that might be introduced by tethering the molecule to a glass surface.
How do you distinguish between stable folds and misfolded kinetic traps?
Kinetic traps should appear as distinct, long-lived populations in FRET histograms. By introducing chaperone proteins, you can study whether this misfolded peak persists or transitions back to the native, unfolded state and into the functional conformation.
Can I study the influence of RNA modifications (e.g., m6A, pseudouridine) on structure?
Site-specific modifications can subtly shift the stability or flexibility of an RNA backbone. Because smFRET is sensitive to sub-nanometer distance changes, you can compare the structural fingerprint of a modified RNA against its unmodified counterpart to see exactly how epitranscriptomic marks alter the molecule’s 3D architecture.
Is it possible to monitor the assembly of large Ribonucleoprotein (RNP) complexes?
FCCS is ideal here – by labelling with two colours, the assembly and stoichiometry of RNP complexes can be quantified. FCS and smFRET can also be used. smFRET is commonly used to study structural dynamics of RNPs, while FCS is suitable for studying binding if the change in mass is sufficient.
Can the system handle highly flexible RNA regions?
For RNAs that have high flexibility and structural heterogeneity, like certain lncRNAs, FCS and smFRET can measure the hydrodynamic radius and conformational fluctuations, providing information on how compact or expanded the RNA is, even if it doesn’t have a fixed 3D shape.
“What sets the EI-FLEX apart is its versatility - it's our go-to for probing kinetics and conformational changes that the other systems can't resolve. This opens up a range of research for us that was previously impossible.”
Professor Edwin Antony, Saint Louis University






