This application note was produced in collaboration with the Robb Lab at the University of Warwick.
In this application note, we demonstrate how Groves et al. performed both single-molecule Förster Resonance Energy Transfer (smFRET) and fluorescence correlation spectroscopy (FCS) on the EI-FLEX, investigating the mechanism of action of two viral SARS-CoV-2 RNA-dependent RNA polymerase (RdRp) inhibitors: a nucleotide competitive inhibitor (remdesivir) and a non-nucleotide inhibitor (sumarin). They designed a doubly labelled RNA hairpin that would act as a reporter for RNA extension by RdRp when analysed using smFRET.
Overview of this application note:
- smFRET can distinguish non-extended from extended RNA species and the effect of RNA-dependent RNA polymerase inhibitors on this process
- FCS provides further context in the form of identifying RdRp stalling on RNA
- These techniques provide complementary data alongside gel-based assays, distinguishing the mechanisms of action for a nucleotide competitive inhibitor and a non-nucleotide inhibitor
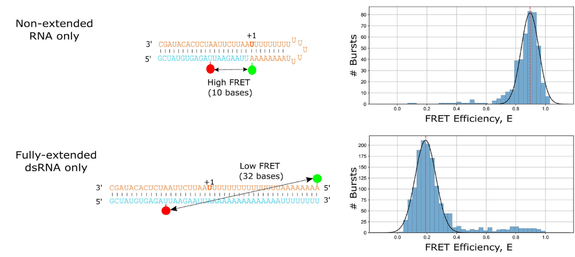
Figure 1 – A doubly-labelled RNA hairpin model captures RNA extension via smFRET
Non-extended RNA produces a high-FRET signal, while fully-extended RNA produces a low-FRET signal. This model permits the investigation of the influence of viral polymerase inhibitors on RNA extension by smFRET.



