In this application note, we demonstrate the use of the EI-FLEX system to recover rates of molecular conformational dynamics, using DNA hairpins as a well-characterised test system. DNA hairpins reversibly transition between open and closed states on a millisecond timescale, with kinetics that are sensitive to NaCl concentration. This makes them an ideal model for validating kinetic measurements using single-molecule Förster Resonance Energy Transfer (smFRET). Our results highlight the utility of the EI-FLEX system in resolving fast dynamic events and underscore its potential as a powerful tool for probing biomolecular mechanisms in solution.
Overview of this application note:
- DNA hairpins are used as a model system to demonstrate how smFRET can capture conformation dynamics for molecular structures
- Burst variance analysis separates static, heterogeneous populations from those that are dynamically undergoing conformational changes
- The influence of salt concentrations on the opening and closing rates of DNA hairpins can be determined by photon-by-photon hidden Markov modelling
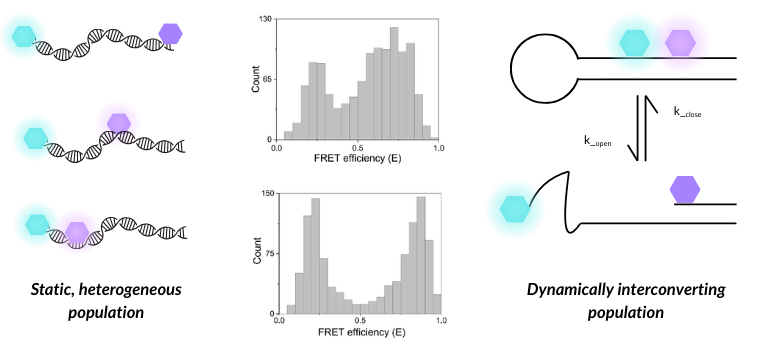
Figure 1 – Distinguishing static, heterogeneous populations from dynamically interconverting biomolecules
A static, heterogeneous population (DNA duplexes with different dye pair placements) can produce similar smFRET histograms to a dynamically interconverting population (DNA hairpin), highlighting the importance of additional dynamic analysis of smFRET data.



