There are no average molecules
In traditional drug discovery, we have long been settling for the average. Whether it’s a plate-based absorbance assay or a Western blot, we look at the collective behaviour of billions of molecules and rely on the mean value to tell the whole story.
However, in biological systems, molecules act as individuals. They move and breathe, shifting between multiple states in response to their environment or binding events. Some states are active, others extremely transient despite being crucial intermediates. Critically, some are precursors to failure that often go unnoticed in the crowd. For the next generation of medicines – from targeted protein degraders (TPDs) to biologics – the average is no longer sufficient to capture the true complexity of these biomolecules.
To bring these compounds to the clinic, we need to stop looking at the crowd and start looking at the individual, failing fast (and crucially, understanding why) to identify the most promising candidates amidst the noise.
Welcome to the era of single-molecule clarity.
In this article, we discuss:
- Introduction to three single-molecule techniques (smFRET, FCS and FCCS)
- Emerging use cases of these technologies in the drug discovery process:
- Conquering undruggable proteins
- Targeted protein degradation of IDPs
- Adding dynamics to cryo-EM structures
- Bringing single-molecule insights into the ‘Lab-in-the-Loop’
A trio of single-molecule tools
Historically, single-molecule biophysics has stayed firmly in specialist labs, with dedicated darkrooms and highly experienced users. Now, with the advent of democratised access to single-molecule spectroscopy tools, integrating these tools into drug discovery pipelines has never been more accessible.
Below, we introduce three single-molecule techniques that offer distinct benefits and insights, while working synergistically with each other to provide a practical toolkit for modern drug development. These techniques are suitable for characterising molecules in complex biofluids, such as serum or cell lysates; performing precise, biophysical measurements in physiologically relevant conditions provides invaluable insights into how a compound, or its target, will behave in vivo.
smFRET: The molecular ruler
Single-Molecule Förster Resonance Energy Transfer (smFRET) is used to measure atomic-scale distances. By labelling two sites on a biomolecule with fluorescent dyes, the distance between them (and how this alters between conformational states) can be measured.
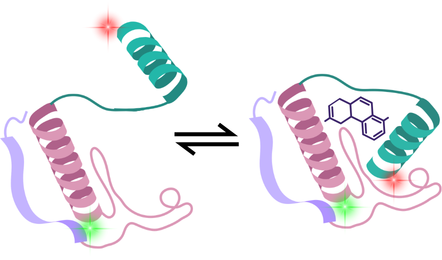
This reveals the dynamic structural landscape of proteins and how small molecules or other compounds influence this. If a drug is designed to lock a receptor in a particular state, smFRET shows whether this is achieved, and most importantly, how many molecules in the entire population are in this expected conformation.
FCS: In-solution binding affinities and aggregation analysis
Fluorescence Correlation Spectroscopy (FCS) measures the diffusion of individual molecules through a femtolitre-scale volume. Given that smaller particles diffuse faster than larger ones or those bound to other biomolecules, size, binding affinity, and concentration can all be calculated from this data.
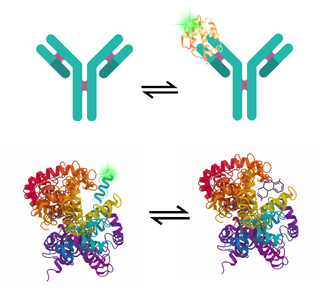
Beyond simple binding, it is a powerful tool for probe competition screening. By using a small, fluorescently labelled probe with a known affinity for the target, FCS can be used to screen libraries of unlabelled compounds to identify candidates that displace the probe. This allows for high-throughput characterisation of native drug candidates without the need for direct labelling. Additionally, FCS also functions as the ultimate early-warning system for aggregation. It can detect the very first seeds of clumping in a biologic formulation, catching these issues long before they cause failures later down the pipeline.
FCCS: Dual-colour molecular co-trajectories
Fluorescence Cross-Correlation Spectroscopy (FCCS) takes FCS a step further. It uses two different-coloured lasers to determine whether two biomolecules (such as a drug and its target) are moving through the solution as a single unit.
This provides a direct, solution-based proof of co-binding, making it ideal for understanding the stoichiometry of complexes, quantifying protein-protein interactions.
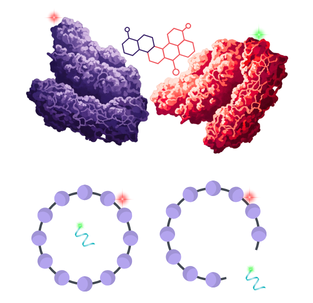
A particularly valuable usecase is the direct confirmation that ternary complexes, such as PROTACs and bispecific antibodies, are interacting with their intended targets. It can also be used for confirming the successful loading of therapeutic cargos for nanoscale drug delivery or detecting when leakage has occurred.
Emerging use cases for single-molecule drug discovery
Conquering ‘undruggable’ targets
For decades, a vast category of proteins has been considered undruggable. Intrinsically disordered proteins (IDPs) don’t have a fixed structure, meaning there is no rigid pocket for a drug to sit in. Given that they exist as a vast ensemble of different conformations, this makes them nearly invisible to traditional structural biology.
This is where single-molecule techniques excel:
- Conformational ensembles via smFRET: While a bulk assay only sees an average of a protein’s shape, smFRET maps the distribution of an IDP’s configurations and is highly synergistic when combined with techniques such as molecule dynamics modelling, SAXS and NMR to generate conformational ensembles for these proteins1.
- Hydrodynamic Profiling via FCS: IDPs often act as hubs for aggregation or multi-protein signalling. FCS measures the hydrodynamic radius of these proteins in real-time2, revealing how environmental factors affect folding, while also detecting if proteins are starting to misfold or clump.
Targeted protein degradation of IDPs
The use of TDPs, such as PROTACs or molecular glues to target IDPs, is increasing, given that they don’t need to stay bound to a specific active site; instead, their role is to bring the IDP into proximity with an E3 ligase, promoting its degradation.
A recent example demonstrates how bulk FRET measurements can be used to characterise a model system where BRD4 is degraded by dBET1 via recruitment to the CRL4CRBN E3 ubiquitin ligase complex3. While high-throughput bulk FRET assays such as this provide methods for measuring ubiquitination kinetics in 384-well plates, single-molecule techniques like smFRET and FCCS take this a step further by resolving the individual binding events and conformational heterogeneity that even the best ensemble assays average out:
| Goal | Technique | The single-molecule advantage |
| Quantifying ternary complex formation | FCCS | Provides absolute proof that a TPD has successfully bridged the IDP and E3 ligase. It confirms binding in solution even if the IDP remains highly disordered. |
| Optimising productive orientation | smFRET | Distinguishes between simple binding and productive binding. It reveals if the IDP and ligase are held in the specific spatial orientation required for degradation to proceed. |
| Confirming target ubiquitination | FCCS | Goes beyond binding to track activity. By detecting the cross-correlation of ubiquitin with the IDP, it quantifies the extent of ubiquitination. |
For more information on how FCCS can quantify ternary complex formation, see our application note here.
Adding dynamics to Cryo-EM structures
Cryo-EM has revolutionised biopharma pipelines by providing atomic-level blueprints of massive protein complexes. However, cryo-EM requires samples to be flash-frozen. This vitrification captures the molecule in a single moment in time, often a state that is chemically trapped or energetically favoured by the freezing process. Capturing the dynamic context of these conformations, and whether any intermediate states exist between them is vital for understanding how these proteins will behave in the body.
smFRET provides this dynamic layer to the static structures identified by cryo-EM. The true heterogeneity of biomolecular samples can be resolved, even if these states are transient or present at low abundance. As discussed above, this is particularly useful for IDPs, or proteins with highly flexible loops that can be tricky to analyse with traditional structural biology techniques.
Further analysis can also reveal whether these molecules are simply individual, static species or whether they are dynamically interconverting (and how the rate of these interconversions is influenced by environmental factors).
Combining the atomic precision of cryo-EM with the single-molecule motion of smFRET therefore provides invaluable insights into why a candidate may be worth taking forward and, perhaps most crucially, why a certain formulation or compound may not be functioning as expected.
Read more on this topic in another article – From atomic detail to molecular motion: Adding dynamics to structural biology.
Bringing single-molecule insights into the
‘Lab-in-the-Loop’
As biopharma moves toward autonomous R&D, accurately validating AI-designed molecules becomes crucial, particularly as Lab-in-the-Loop systems are deployed. Single-molecule data is hugely valuable for supporting the development of these models, as it provides dynamic information and captures the true heterogeneity that exists within nature.
Why single-molecule data is crucial for training Lab-in-the-Loop systems:
- Most AI models (like AlphaFold-3) were trained on static snapshots. To design the next generation of therapeutics, AI needs to understand dynamics. Single-molecule data provides the ground truth of how a protein folds and moves, complementing static structural information, while also capturing binding information and aggregation tendencies.
- Bulk assays are often noisy because they average out populations, such as those in rare conformational states, that may make a drug successful. Single-molecule techniques capture this heterogeneity, enabling AI models to learn faster with fewer samples.
Multiomics and multimodal data are increasingly being deployed at scale, informing AI and machine learning models to support the development of new drugs and companion diagnostics. The addition of dynamic single-molecule data could be invaluable here, providing the mechanistic context of how a mutation influences the conformational state of a protein and the downstream impact this has on the wider proteome and multiome.
While the integration of single-cell imaging and omics has already been proposed as a powerful tool to resolve cellular heterogeneity, researchers note that significant challenges remain. As highlighted by Watson et al., these include the low-throughput nature of high-resolution imaging and the immense computational demands of merging multidimensional datasets4.
Techniques like smFRET and FCS offer a unique opportunity to alleviate these burdens; by providing high-precision ‘biophysical anchors,’ they allow AI models to prioritise causal pathways over random correlations, effectively acting as a filter to reduce the noise inherent in high-dimensional omics data. By training models on rich, dynamic data now, multimodal diagnostics and drug discovery can be supported by a more robust framework for years to come.
Conclusion
We have entered an era where relying on the average is a risk. In the race to the clinic, the winners are the ones who can see the outliers: the hidden aggregates, the flexible, ephemeral states of IDPs, and the precise stoichiometry of a ternary complex.
By combining the structural precision of Cryo-EM with the dynamic insights of single-molecule techniques and feeding that data into AI-driven loops, new compounds can be designed and validated with confidence, reducing the risk of costly late-stage failures.
Ultimately, we must shift from analysing the crowd and start looking at the individual. Because in biology, there are no average molecules.
Interested in learning more? Click here to explore a world of Exciting insights and applications for single-molecule techniques.
References
- Naudi-Fabra, S., Tengo, M., Jensen, M. R., Blackledge, M. & Milles, S. Quantitative Description of Intrinsically Disordered Proteins Using Single-Molecule FRET, NMR, and SAXS. J. Am. Chem. Soc. 143, 20109–20121 (2021).
- Białobrzewski, M. K. et al. Diversity of hydrodynamic radii of intrinsically disordered proteins. European Biophysics Journal 52, 607–618 (2023).
- Yu, C. H. et al. Single-Assay Characterization of Ternary Complex Assembly and Activity in Targeted Protein Degradation. bioRxiv 2025.08.20.671298 (2025) doi:10.1101/2025.08.20.671298.
- Watson, E. R., Taherian Fard, A. & Mar, J. C. Computational Methods for Single-Cell Imaging and Omics Data Integration. Front. Mol. Biosci. Volume 8-2021, (2022).



